Our most recent work on actin filaments quantification! Oral presentation at ICIP 2020
Presenter: Yi Liu
Our most recent work on actin filaments quantification! Oral presentation at ICIP 2020
Presenter: Yi Liu
---------------------------------------------------------------------------------------------------------------------------------
Demo of stromule tracking on actin/microtubule filaments
tracking start from 45s
---------------------------------------------------------------------------------------------------------------------------------
Demo of Our Instance Segmentation Result
Since there are too many filaments in one image, we present indiviudal filaments one by one
---------------------------------------------------------------------------------------------------------------------------------
Part1: Binary Segmentation Microtubules and Actin filaments
In this part, we introduce a novel deep network architecture for filament segmentation in confocal microscopy images that improves upon the state-of-the-art U-net and SOAX methods. We also propose a strategy for data annotation, and create datasets for microtubule and actin filaments.
Liu Y, Treible W, Kolagunda A, Nedo A, Saponaro P, Caplan J, Kambhamettu C. Densely connected stacked u-network for filament segmentation in microscopy images. InProceedings of the European Conference on Computer Vision (ECCV) 2018 (pp. 0-0).
}
Part2: Instance Segmentation of Microtubules
In this Part, we introduce a orientation-aware neural network, which contains six orientation-associated branches. Each branch detects filaments with specific range of orientations, thus separating them at junctions, and turning intersections to overpasses. A terminus pairing algorithm is also proposed to regroup filaments from different branches, and achieve individual filaments extraction.
Liu, Y., Kolagunda, A., Treible, W., Nedo, A., Caplan, J., & Kambhamettu, C. (2019). Intersection to Overpass: Instance Segmentation on Filamentous Structures With an Orientation-Aware Neural Network and Terminus Pairing Algorithm. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition Workshops (pp. 0-0).
Part3: Instance Segmentation of Actins
In this Part, we adopt a Convolutional Neural Network (CNN)to segment actin filaments first, and then we utilize a mod-ified Resnet to detect junctions and endpoints of filaments.With binary segmentation and detected keypoints, we applya fast marching algorithm to obtain the number and length ofeach actin filament in microscopic images.
Y. Liu, A. Nedo, K. Seward, J. Caplan and C. Kambhamettu, "Quantifying Actin Filaments in Microscopic Images using Keypoint Detection Techniques and A Fast Marching Algorithm," 2020 IEEE International Conference on Image Processing (ICIP), Abu Dhabi, United Arab Emirates, 2020, pp. 2506-2510, doi: 10.1109/ICIP40778.2020.9191337.
Details of Part 1: Binary Segmentation Microtubules and Actin filaments
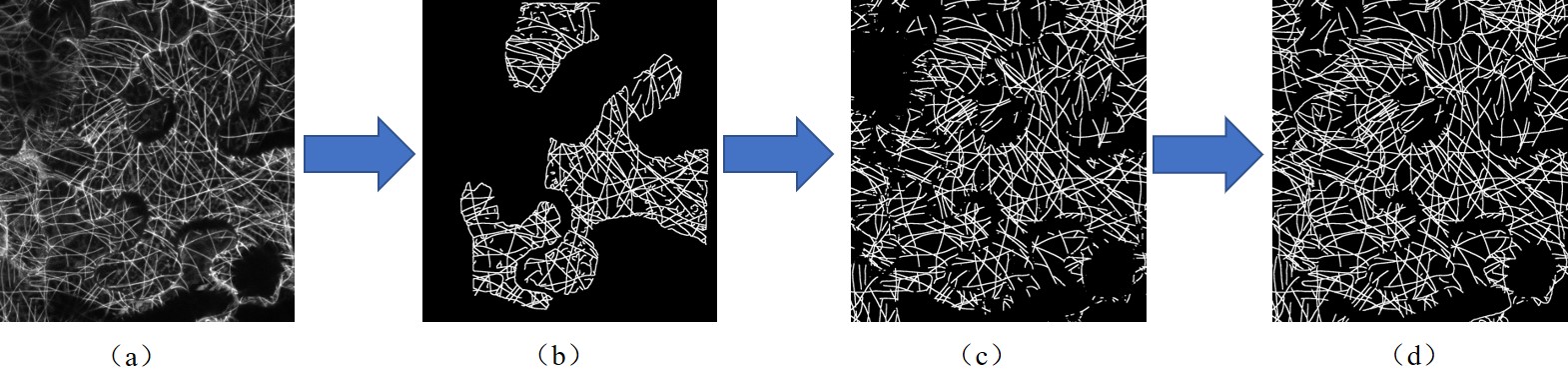
Annotation process. (a) Original image of mircrotubules. (b) SOAX Segmentation result
(c) Segmentation result of U-net which is trained with SOAX segmentation result
(d) Manually labeled ground truth based on U-net result
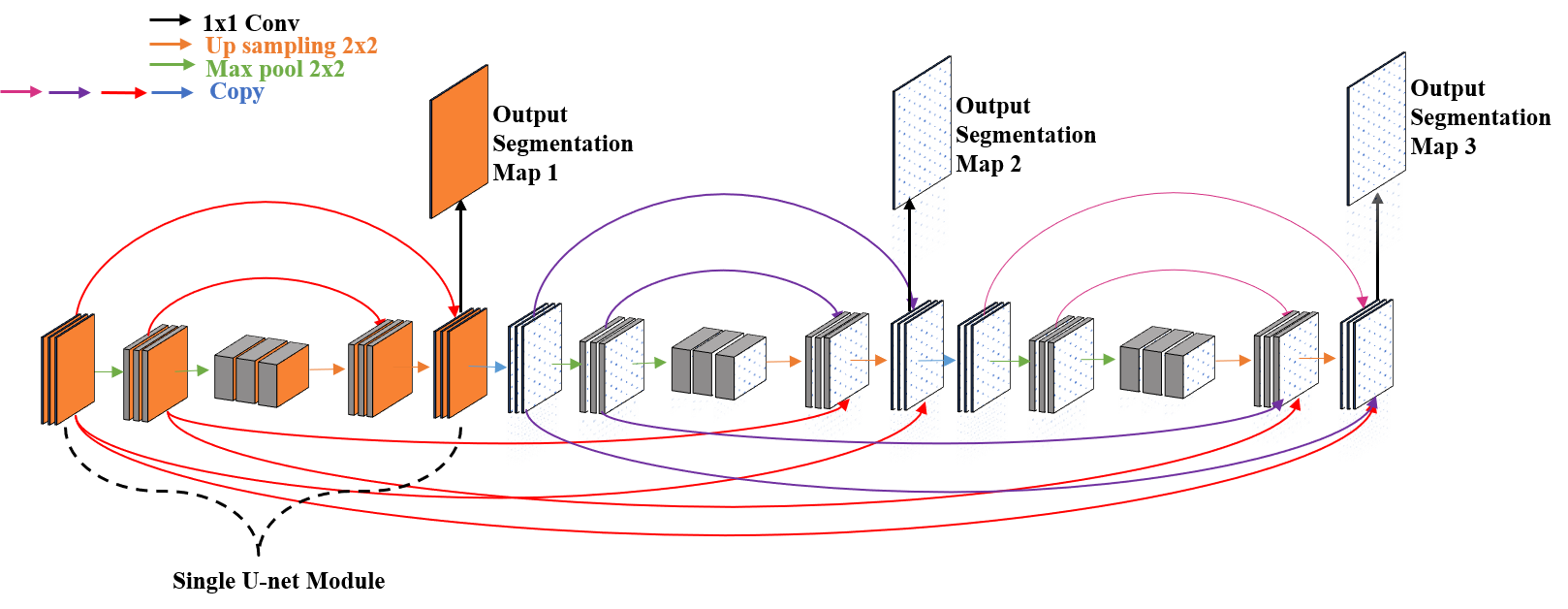
An illustration of our proposed network
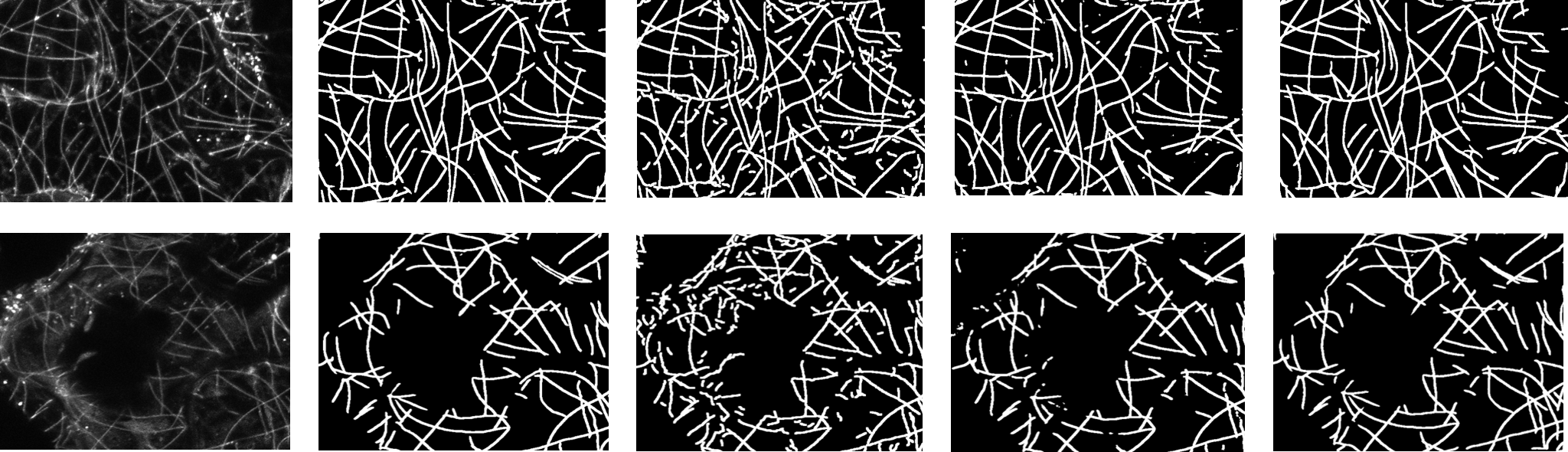

Segmentation of microtubules.
From left to right: original image, ground truth,SOAX, U-net module, proposed network with cross connection
Details of Part 2: Instance Segmentation of Microtubules
Intuition of our proposed method:
Regrouping the fragments at intersections is the bottle neck to group individual filaments.
We propose an orientation-aware neural network which can sort filaments into six bins according to their orientation angle
Each filament will automatically classiffied into certain layer, the intuition is similar like turning an intersection to overpass
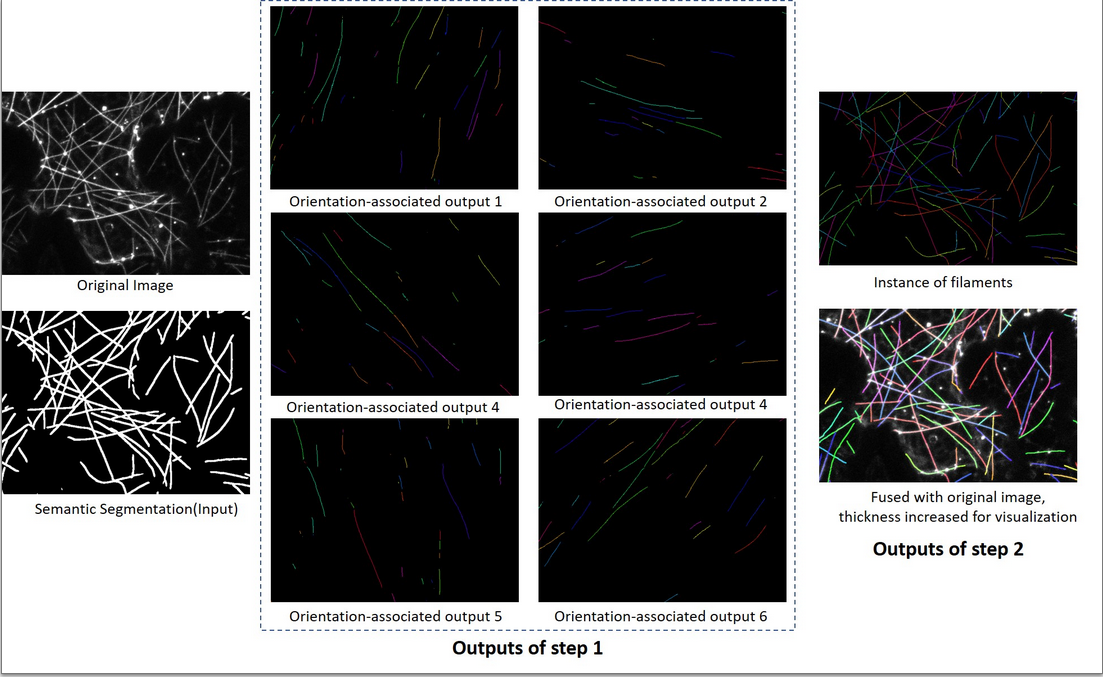

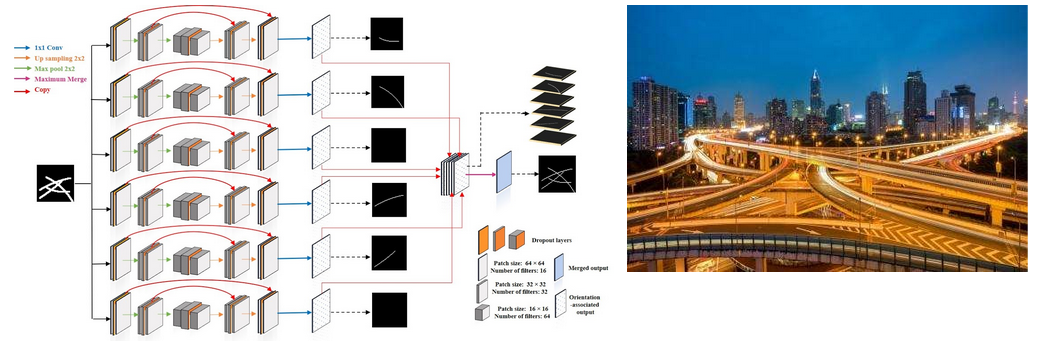
An illustration of our proposed pipeline:
Step 1: The network take a segmentation as input, and six orientation-associated outputs willbe generated.
Step 2: The proposed algorithm is applied on these six outputs and regroup fragments into individual filaments.
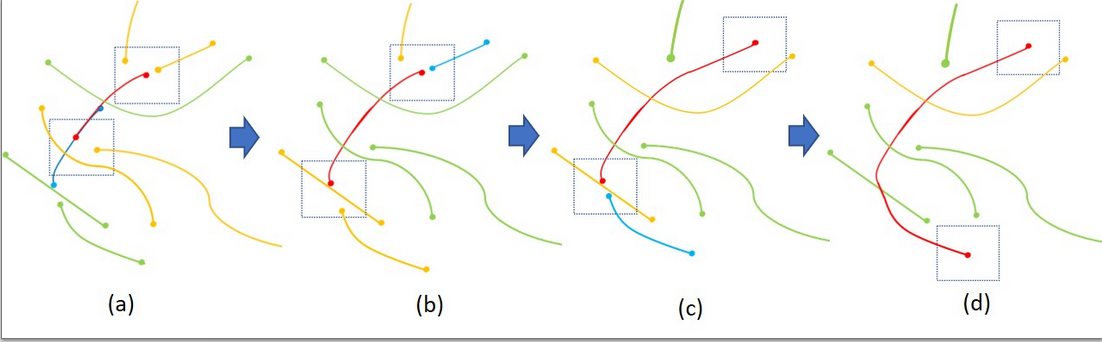
An example of recursively grouping the most eligiblecandidate from different layers
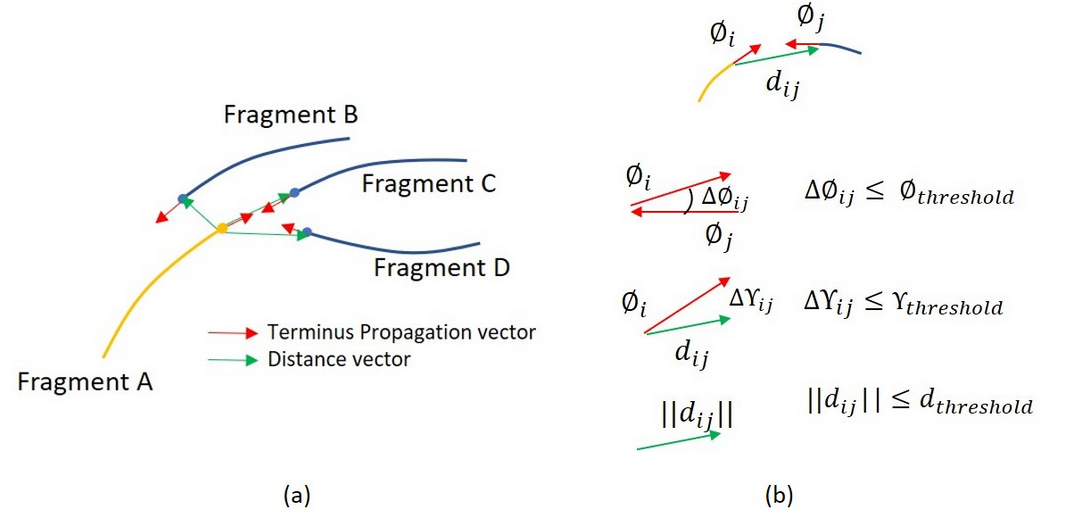
Reconnection by their geometric properties
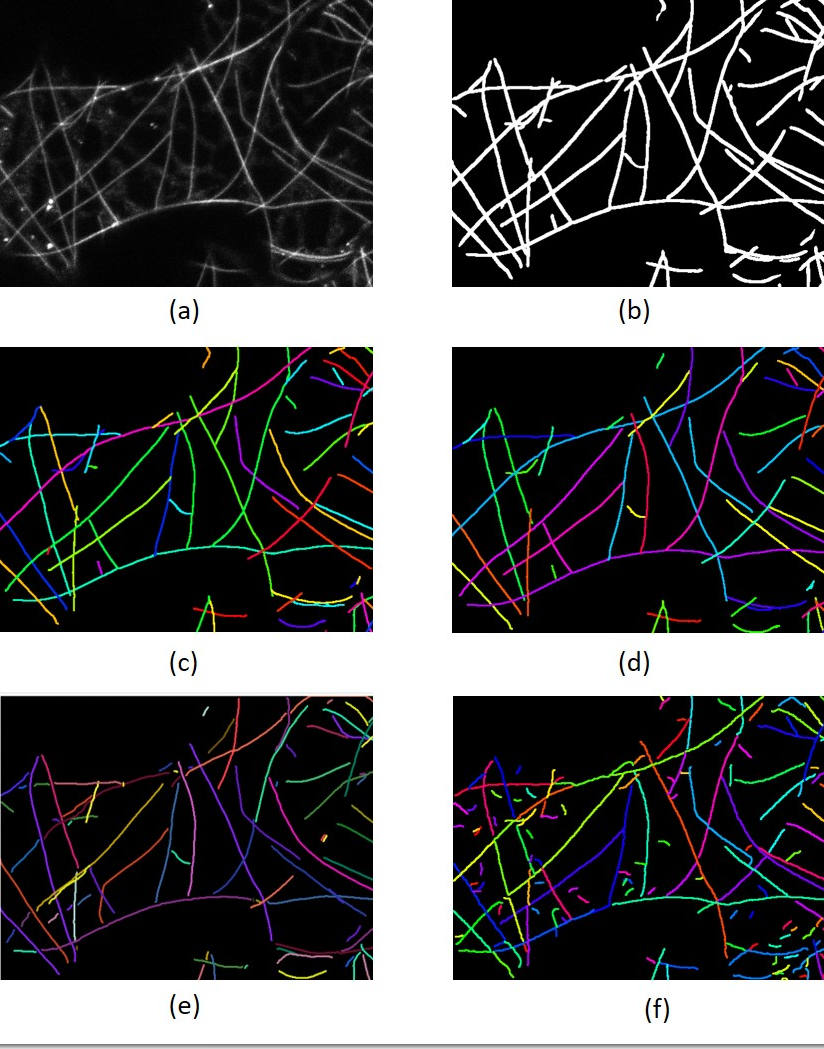

Comparison of different methods on extracting individ-ual filament. (a) Original image (b) Segmentation
(c) Manuallycorrected labels (d) Result of our proposed method (e) SFINE
(f)SOAX without filtering out small fragments. Different colors in-dicate different labels, and some color might look similar, but theyare actually different labels.
Details of Part 3: Instance Segmentation of Actins
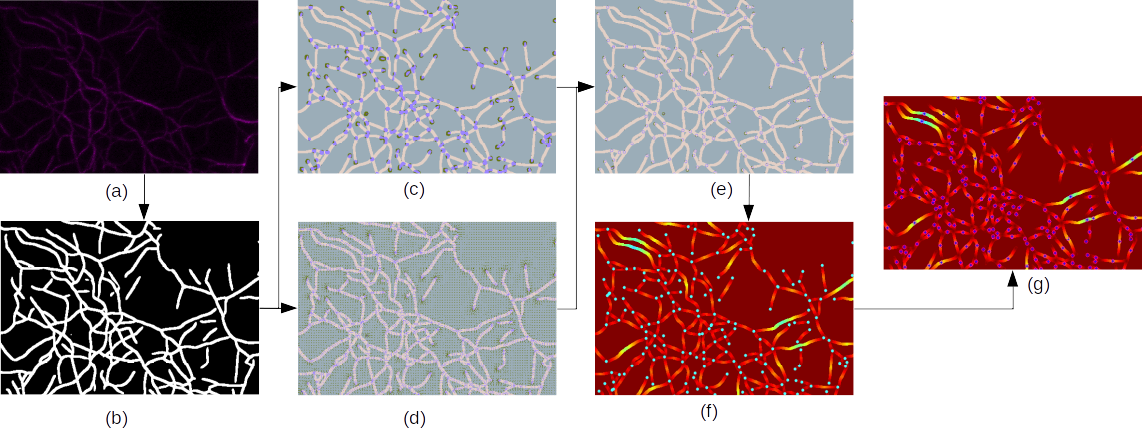

The pipeline of our proposed approach. (a) Microscopic image of actin filaments. (b) Binary segmentation.
(c) Heatmapsof junctions and endpoints. (d) Offset maps of junctions and endpoints. (e) Prediction of junctions and endpoints.
(f) Geodesicdistance map from junctions and endpoints, which are color of cyan.
(g) The local maximum value of the geodesic distancemap, which is represented in blue with a pink circle.
(a) to (b): We use a neural network to obtain binary segmentation.
(b)to (c),(d) and (e): Our modified Resnet takes binary segmentation results as inputs and outputs offset maps and heatmaps ofjunctions and endpoints.
Then we integrate (c) and (d) to obtain refined locations of these points.
(e) to (f) and (g): We setpredicted keypoints as start points and utilize a fast marching algorithm to calculate the geodesic distance map. The local peakvalues of the geodesic distance map can represent the half-length value of the actin filaments.
© Yi Liu, Chandra Kambhamettu, Jeff Caplan and the VIMS Lab. All rights reserved. | Design adapted from TEMPLATED.